NicoScribe (talk | contribs) more specific categorization |
Plantsurfer (talk | contribs) m Replaced VE ref names using RefRenamernamed citatoins |
||
| Line 17: | Line 17: | ||
}} |
}} |
||
'''Asgard''' or '''Asgardarchaeota'''<ref name="pmid28604769">{{cite journal |last=Da Cunha |first=Violette |last2=Gaia |first2=Morgan |last3=Gadelle |first3=Daniele |last4=Nasir |first4=Arshan |last5=Forterre |first5=Patrick |display-authors=3 |title=Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes |journal=PLOS Genetics |volume=13 |issue=6 |pages=e1006810 |date=June 2017 |pmid=28604769 |pmc=5484517 |doi=10.1371/journal.pgen.1006810 }}</ref> is a proposed [[superphylum]] consisting of a group of [[archaea]] that contain eukaryotic signature proteins.<ref name="Zaremba">{{cite journal |last1=Zaremba-Niedzwiedzka |first1=Katarzyna |last2=Caceres |first2=Eva F. |last3=Saw |first3=Jimmy H. |last4=Bäckström |first4=Disa |last5=Juzokaite |first5=Lina |last6=Vancaester |first6=Emmelien |last7=Seitz |first7=Kiley W. |last8=Anantharaman |first8=Karthik |last9=Starnawski |first9=Piotr |display-authors=3 |title=Asgard archaea illuminate the origin of eukaryotic cellular complexity |journal=Nature |volume=541 |issue=7637 |pages=353–358 |date=January 2017 |pmid=28077874 |doi=10.1038/nature21031 |osti=1580084 |s2cid=4458094 |bibcode=2017Natur.541..353Z }}</ref> It appears that the [[eukaryote]]s, the [[Domain (biology)|domain]] that contains the [[animal]]s, [[plant]]s, and [[Fungus|fungi]], [[Eukaryogenesis|emerged within the Asgard]],<ref name="Eme">{{cite journal |last1=Eme |first1=Laura|last2=Spang |first2=Anja |last3=Lombard|first3=Jonathan |last4=Stairs |first4=Courtney W. |last5=Ettema |first5=Thijs J. G. |title=Archaea and the origin of eukaryotes |journal=Nature Reviews. Microbiology |volume=15 |issue=12 |pages=711–723 |date=November 2017 |pmid=29123225 |doi=10.1038/nrmicro.2017.133 |s2cid=8666687 }}</ref> in a branch containing the Heimdallarchaeota.<ref name="Williams 138–147">{{cite journal |last1=Williams |first1=Tom A. |last2=Cox |first2=Cymon J. |last3=Foster |first3=Peter G. |last4=Szöllősi |first4=Gergely J. |last5=Embley |first5=T. Martin |title=Phylogenomics provides robust support for a two-domains tree of life |journal=Nature Ecology & Evolution |volume=4 |issue=1 |pages=138–147 |date=January 2020 |pmid=31819234 |pmc=6942926 |doi=10.1038/s41559-019-1040-x }}</ref> This supports the [[two-domain system]] of classification over the [[three-domain system]].<ref name=" |
'''Asgard''' or '''Asgardarchaeota'''<ref name="pmid28604769">{{cite journal |last=Da Cunha |first=Violette |last2=Gaia |first2=Morgan |last3=Gadelle |first3=Daniele |last4=Nasir |first4=Arshan |last5=Forterre |first5=Patrick |display-authors=3 |title=Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes |journal=PLOS Genetics |volume=13 |issue=6 |pages=e1006810 |date=June 2017 |pmid=28604769 |pmc=5484517 |doi=10.1371/journal.pgen.1006810 }}</ref> is a proposed [[superphylum]] consisting of a group of [[archaea]] that contain eukaryotic signature proteins.<ref name="Zaremba">{{cite journal |last1=Zaremba-Niedzwiedzka |first1=Katarzyna |last2=Caceres |first2=Eva F. |last3=Saw |first3=Jimmy H. |last4=Bäckström |first4=Disa |last5=Juzokaite |first5=Lina |last6=Vancaester |first6=Emmelien |last7=Seitz |first7=Kiley W. |last8=Anantharaman |first8=Karthik |last9=Starnawski |first9=Piotr |display-authors=3 |title=Asgard archaea illuminate the origin of eukaryotic cellular complexity |journal=Nature |volume=541 |issue=7637 |pages=353–358 |date=January 2017 |pmid=28077874 |doi=10.1038/nature21031 |osti=1580084 |s2cid=4458094 |bibcode=2017Natur.541..353Z }}</ref> It appears that the [[eukaryote]]s, the [[Domain (biology)|domain]] that contains the [[animal]]s, [[plant]]s, and [[Fungus|fungi]], [[Eukaryogenesis|emerged within the Asgard]],<ref name="Eme">{{cite journal |last1=Eme |first1=Laura|last2=Spang |first2=Anja |last3=Lombard|first3=Jonathan |last4=Stairs |first4=Courtney W. |last5=Ettema |first5=Thijs J. G. |title=Archaea and the origin of eukaryotes |journal=Nature Reviews. Microbiology |volume=15 |issue=12 |pages=711–723 |date=November 2017 |pmid=29123225 |doi=10.1038/nrmicro.2017.133 |s2cid=8666687 }}</ref> in a branch containing the Heimdallarchaeota.<ref name="Williams 138–147">{{cite journal |last1=Williams |first1=Tom A. |last2=Cox |first2=Cymon J. |last3=Foster |first3=Peter G. |last4=Szöllősi |first4=Gergely J. |last5=Embley |first5=T. Martin |title=Phylogenomics provides robust support for a two-domains tree of life |journal=Nature Ecology & Evolution |volume=4 |issue=1 |pages=138–147 |date=January 2020 |pmid=31819234 |pmc=6942926 |doi=10.1038/s41559-019-1040-x }}</ref> This supports the [[two-domain system]] of classification over the [[three-domain system]].<ref name="Nobs-2022">{{cite journal |last1=Nobs |first1=Stephanie-Jane |last2=MacLeod |first2=Fraser I. |last3=Wong |first3=Hon Lun |last4=Burns |first4=Brendan P. |title=Eukarya the chimera: eukaryotes, a secondary innovation of the two domains of life? |journal=Trends in Microbiology |volume=30 |issue=5 |pages=421–431 |date=May 2022 |pmid=34863611 |doi=10.1016/j.tim.2021.11.003 |s2cid=244823103 }}</ref><ref name="Doolittle-2020">{{cite journal |last=Doolittle |first=W. Ford |title=Evolution: Two Domains of Life or Three? |journal=Current Biology |volume=30 |issue=4 |pages=R177–R179 |date=February 2020 |pmid=32097647 |doi=10.1016/j.cub.2020.01.010 |doi-access=free }}</ref> |
||
==Discovery and nomenclature== |
==Discovery and nomenclature== |
||
| Line 33: | Line 33: | ||
Asgard members encode many eukaryotic signature proteins, including novel [[Guanosine triphosphate|GTPases]], membrane-remodelling proteins like [[ESCRT]] and [[CHMP4B|SNF7]], a [[ubiquitin]] modifier system, and [[N-glycosylation]] pathway homologs.<ref name="Zaremba"/> |
Asgard members encode many eukaryotic signature proteins, including novel [[Guanosine triphosphate|GTPases]], membrane-remodelling proteins like [[ESCRT]] and [[CHMP4B|SNF7]], a [[ubiquitin]] modifier system, and [[N-glycosylation]] pathway homologs.<ref name="Zaremba"/> |
||
Asgard archaeons have a regulated [[actin]] [[cytoskeleton]], and the [[profilin]]s and [[gelsolin]]s they use can interact with eukaryotic actins.<ref>{{cite journal |last=Akıl |first=Caner |last2=Robinson |first2=Robert C. |title=Genomes of Asgard archaea encode profilins that regulate actin |journal=Nature |volume=562 |issue=7727 |pages=439–443 |date=October 2018 |pmid=30283132 |doi=10.1038/s41586-018-0548-6 |s2cid=52917038 |bibcode=2018Natur.562..439A }}</ref><ref>{{cite journal |last=Akıl |first=Caner |last2=Tran |first2=Linh T. |last3=Orhant-Prioux |first3=Magali |last4=Baskaran |first4=Yohendran |last5=Manser |first5=Edward |last6=Blanchoin |first6=Laurent |last7=Robinson |first7=Robert C. |title=Insights into the evolution of regulated actin dynamics via characterization of primitive gelsolin/cofilin proteins from Asgard archaea |journal=Proceedings of the National Academy of Sciences of the United States of America |volume=117 |issue=33 |pages=19904–19913 |date=August 2020 |pmid=32747565 |pmc=7444086 |doi=10.1073/pnas.2009167117 |doi-access=free |biorxiv=10.1101/768580 }}</ref> In addition, Asgard archaea [[tubulin]] from hydrothermal-living Odinarchaeota (OdinTubulin) was identified as a genuine tubulin. OdinTubulin forms protomers and protofilaments most similar to eukaryotic microtubules, yet assembles into ring systems more similar to [[FtsZ]], indicating that OdinTubulin may represent an evolution intermediate between FtsZ and [[microtubule]]-forming tubulins.<ref>{{cite journal |last1=Akıl |first1=Caner |last2=Ali |first2=Samson |last3=Tran |first3=Linh T. |last4=Gaillard |first4=Jeremie |last5=Li |first5=Wenfei |last6=Hayashida |first6=Kenichi |last7=Mika |first7=Hirose |last8=Oshima |first8=Atsunori |last9=Fujishima |first9=Kosuke |last10=Blanchoin |first10=Laurent |last11=Narita |first11=Akihiro |last12=Robinson |first12=Robert C. |display-authors=3 |title=Structure and dynamics of Odinarchaeota tubulin and the implications for eukaryotic microtubule evolution |journal=Science Advances |volume=8 |issue=12 |pages=eabm2225 |date=March 2022 |pmid=35333570 |pmc=8956254 |doi=10.1126/sciadv.abm2225 |bibcode=2022SciA....8M2225A }}</ref> They also seem to form vesicles under [[cryogenic electron microscopy]]. Some may have a [[PKD domain]] [[S-layer]].<ref name=" |
Asgard archaeons have a regulated [[actin]] [[cytoskeleton]], and the [[profilin]]s and [[gelsolin]]s they use can interact with eukaryotic actins.<ref>{{cite journal |last=Akıl |first=Caner |last2=Robinson |first2=Robert C. |title=Genomes of Asgard archaea encode profilins that regulate actin |journal=Nature |volume=562 |issue=7727 |pages=439–443 |date=October 2018 |pmid=30283132 |doi=10.1038/s41586-018-0548-6 |s2cid=52917038 |bibcode=2018Natur.562..439A }}</ref><ref>{{cite journal |last=Akıl |first=Caner |last2=Tran |first2=Linh T. |last3=Orhant-Prioux |first3=Magali |last4=Baskaran |first4=Yohendran |last5=Manser |first5=Edward |last6=Blanchoin |first6=Laurent |last7=Robinson |first7=Robert C. |title=Insights into the evolution of regulated actin dynamics via characterization of primitive gelsolin/cofilin proteins from Asgard archaea |journal=Proceedings of the National Academy of Sciences of the United States of America |volume=117 |issue=33 |pages=19904–19913 |date=August 2020 |pmid=32747565 |pmc=7444086 |doi=10.1073/pnas.2009167117 |doi-access=free |biorxiv=10.1101/768580 }}</ref> In addition, Asgard archaea [[tubulin]] from hydrothermal-living Odinarchaeota (OdinTubulin) was identified as a genuine tubulin. OdinTubulin forms protomers and protofilaments most similar to eukaryotic microtubules, yet assembles into ring systems more similar to [[FtsZ]], indicating that OdinTubulin may represent an evolution intermediate between FtsZ and [[microtubule]]-forming tubulins.<ref>{{cite journal |last1=Akıl |first1=Caner |last2=Ali |first2=Samson |last3=Tran |first3=Linh T. |last4=Gaillard |first4=Jeremie |last5=Li |first5=Wenfei |last6=Hayashida |first6=Kenichi |last7=Mika |first7=Hirose |last8=Oshima |first8=Atsunori |last9=Fujishima |first9=Kosuke |last10=Blanchoin |first10=Laurent |last11=Narita |first11=Akihiro |last12=Robinson |first12=Robert C. |display-authors=3 |title=Structure and dynamics of Odinarchaeota tubulin and the implications for eukaryotic microtubule evolution |journal=Science Advances |volume=8 |issue=12 |pages=eabm2225 |date=March 2022 |pmid=35333570 |pmc=8956254 |doi=10.1126/sciadv.abm2225 |bibcode=2022SciA....8M2225A }}</ref> They also seem to form vesicles under [[cryogenic electron microscopy]]. Some may have a [[PKD domain]] [[S-layer]].<ref name="Imachi-2020">{{cite journal |last1=Imachi |first1=Hiroyuki |last2=Nobu |first2=Masaru K. |last3=Nakahara |first3=Nozomi|last4=Morono |first4=Yuki |last5=Ogawara |first5=Miyuki |last6=Takaki |first6=Yoshihiro |last7=Takano |first7=Yoshinori |last8=Uematsu |first8=Katsuyuki |last9=Ikuta |first9=Tetsuro |last10=Ito |first10=Motoo |last11=Matsui |first11=Yohei |display-authors=3 |title=Isolation of an archaeon at the prokaryote-eukaryote interface |journal=Nature |volume=577 |issue=7791 |pages=519–525 |date=January 2020 |pmid=31942073 |pmc=7015854 |doi=10.1038/s41586-019-1916-6 |bibcode=2020Natur.577..519I }}</ref> They also share the three-way ES39 expansion in [[LSU rRNA]] with eukaryotes.<ref>{{cite journal |last1=Penev |first1=Petar I. |last2=Fakhretaha-Aval |first2=Sara |last3=Patel |first3=Vaishnavi J. |last4=Cannone |first4=Jamie J. |last5=Gutell |first5=Robin R. |last6=Petrov |first6=Anton S. |last7=Williams |first7=Loren Dean |last8=Glass |first8=Jennifer B. |display-authors=3 |title=Supersized Ribosomal RNA Expansion Segments in Asgard Archaea |journal=Genome Biology and Evolution |volume=12 |issue=10 |pages=1694–1710 |date=October 2020 |pmid=32785681 |pmc=7594248 |doi=10.1093/gbe/evaa170 |doi-access=free }}</ref> |
||
===Metabolism=== |
===Metabolism=== |
||
| Line 41: | Line 41: | ||
</gallery> |
</gallery> |
||
Asgard archaea are generally [[obligate anaerobe]]s, though Kariarchaeota, Gerdarchaeota and Hodarchaeota may be [[facultative aerobes]].<ref name="Liu_2020">{{cite journal |last=Liu |first=Yang |last2=Makarova |first2=Kira S. |last3=Huang |first3=Wen-Cong |last4=Wolf |first4=Yuri I. |last5=Nikolskaya |first5=Anastasia |last6=Zhang |first6=Xinxu |last7=Cai |first7=Mingwei |last8=Zhang |first8=Cui-Jing |last9=Xu |first9=Wei |last10=Luo |first10=Zhuhua |last11=Cheng |first11=Lei |last12=Koonin |first12=Eugene V. |author12-link=Eugene Koonin |last13=Li |first13=Meng |display-authors=3 |title=Expanding diversity of Asgard archaea and the elusive ancestry of eukaryotes |journal=bioRxiv |year=2020 |doi=10.1101/2020.10.19.343400 |s2cid=225056970 |url=https://www.biorxiv.org/content/10.1101/2020.10.19.343400v3.full}}</ref> They have a [[Wood–Ljungdahl pathway]] and perform [[glycolysis]]. Members can be [[autotroph]]s, [[heterotroph]]s, or [[phototroph]]s using [[heliorhodopsin]].<ref name="kindler2019">{{cite journal |last1=MacLeod |first1=Fraser |last2=Kindler |first2=Gareth S. |last3=Wong |first3=Hon Lun |last4=Chen |first4=Ray |last5=Burns |first5=Brendan P. |title=Asgard archaea: Diversity, function, and evolutionary implications in a range of microbiomes |journal=AIMS Microbiology |volume=5 |issue=1 |pages=48–61 |date=2019 |pmid=31384702 |pmc=6646929 |doi=10.3934/microbiol.2019.1.48 }}</ref> One member, [[Candidatus Prometheoarchaeum syntrophicum|''Candidatus'' ''Prometheoarchaeum syntrophicum'']], is [[syntrophy|syntrophic]] with a sulfur-reducing proteobacteria and a [[methanogenic]] archaea.<ref name=" |
Asgard archaea are generally [[obligate anaerobe]]s, though Kariarchaeota, Gerdarchaeota and Hodarchaeota may be [[facultative aerobes]].<ref name="Liu_2020">{{cite journal |last=Liu |first=Yang |last2=Makarova |first2=Kira S. |last3=Huang |first3=Wen-Cong |last4=Wolf |first4=Yuri I. |last5=Nikolskaya |first5=Anastasia |last6=Zhang |first6=Xinxu |last7=Cai |first7=Mingwei |last8=Zhang |first8=Cui-Jing |last9=Xu |first9=Wei |last10=Luo |first10=Zhuhua |last11=Cheng |first11=Lei |last12=Koonin |first12=Eugene V. |author12-link=Eugene Koonin |last13=Li |first13=Meng |display-authors=3 |title=Expanding diversity of Asgard archaea and the elusive ancestry of eukaryotes |journal=bioRxiv |year=2020 |doi=10.1101/2020.10.19.343400 |s2cid=225056970 |url=https://www.biorxiv.org/content/10.1101/2020.10.19.343400v3.full}}</ref> They have a [[Wood–Ljungdahl pathway]] and perform [[glycolysis]]. Members can be [[autotroph]]s, [[heterotroph]]s, or [[phototroph]]s using [[heliorhodopsin]].<ref name="kindler2019">{{cite journal |last1=MacLeod |first1=Fraser |last2=Kindler |first2=Gareth S. |last3=Wong |first3=Hon Lun |last4=Chen |first4=Ray |last5=Burns |first5=Brendan P. |title=Asgard archaea: Diversity, function, and evolutionary implications in a range of microbiomes |journal=AIMS Microbiology |volume=5 |issue=1 |pages=48–61 |date=2019 |pmid=31384702 |pmc=6646929 |doi=10.3934/microbiol.2019.1.48 }}</ref> One member, [[Candidatus Prometheoarchaeum syntrophicum|''Candidatus'' ''Prometheoarchaeum syntrophicum'']], is [[syntrophy|syntrophic]] with a sulfur-reducing proteobacteria and a [[methanogenic]] archaea.<ref name="Imachi-2020" /> |
||
The [[RuBisCO]] they have is not carbon-fixing, but likely used for nucleoside salvaging.<ref name="kindler2019"/> |
The [[RuBisCO]] they have is not carbon-fixing, but likely used for nucleoside salvaging.<ref name="kindler2019"/> |
||
| Line 53: | Line 53: | ||
The phylum Heimdallarchaeota was found in 2017 to have N-terminal core [[Histone code|histone tails]], a feature previously thought to be exclusively eukaryotic. Two other archaeal phyla, both outside of Asgard, were found to also have tails in 2018.<ref name=pmid30212449>{{cite journal |last=Henneman |first=Bram |last2=van Emmerik |first2=Clara |last3=van Ingen |first3=Hugo |last4=Dame |first4=Remus T. |title=Structure and function of archaeal histones |journal=PLOS Genetics |volume=14 |issue=9 |pages=e1007582 |date=September 2018 |pmid=30212449 |pmc=6136690 |doi=10.1371/journal.pgen.1007582 |bibcode=2018BpJ...114..446H }}</ref> |
The phylum Heimdallarchaeota was found in 2017 to have N-terminal core [[Histone code|histone tails]], a feature previously thought to be exclusively eukaryotic. Two other archaeal phyla, both outside of Asgard, were found to also have tails in 2018.<ref name=pmid30212449>{{cite journal |last=Henneman |first=Bram |last2=van Emmerik |first2=Clara |last3=van Ingen |first3=Hugo |last4=Dame |first4=Remus T. |title=Structure and function of archaeal histones |journal=PLOS Genetics |volume=14 |issue=9 |pages=e1007582 |date=September 2018 |pmid=30212449 |pmc=6136690 |doi=10.1371/journal.pgen.1007582 |bibcode=2018BpJ...114..446H }}</ref> |
||
In January 2020, scientists found ''Candidatus'' ''Prometheoarchaeum syntrophicum'', a member of the Lokiarcheota, engaging in [[cross-feeding]] with two bacterial species. Drawing an analogy to [[symbiogenesis]], they consider this relationship a possible link between the simple [[prokaryotic]] microorganisms and the complex [[eukaryotic]] microorganisms occurring approximately two billion years ago.<ref name="NYT-2020115">{{cite news |last=Zimmer |first=Carl |author-link=Carl Zimmer |title=This Strange Microbe May Mark One of Life's Great Leaps - A organism living in ocean muck offers clues to the origins of the complex cells of all animals and plants. |url=https://www.nytimes.com/2020/01/15/science/cells-eukaryotes-archaea.html |date=15 January 2020 |work=[[The New York Times]] |access-date=16 January 2020 }}</ref><ref name=" |
In January 2020, scientists found ''Candidatus'' ''Prometheoarchaeum syntrophicum'', a member of the Lokiarcheota, engaging in [[cross-feeding]] with two bacterial species. Drawing an analogy to [[symbiogenesis]], they consider this relationship a possible link between the simple [[prokaryotic]] microorganisms and the complex [[eukaryotic]] microorganisms occurring approximately two billion years ago.<ref name="NYT-2020115">{{cite news |last=Zimmer |first=Carl |author-link=Carl Zimmer |title=This Strange Microbe May Mark One of Life's Great Leaps - A organism living in ocean muck offers clues to the origins of the complex cells of all animals and plants. |url=https://www.nytimes.com/2020/01/15/science/cells-eukaryotes-archaea.html |date=15 January 2020 |work=[[The New York Times]] |access-date=16 January 2020 }}</ref><ref name="Imachi-2020"/> |
||
== Classification == |
== Classification == |
||
Revision as of 21:30, 6 June 2023
| Asgard | |
|---|---|
| Scientific classification | |
| Domain: | Archaea |
| Kingdom: | Proteoarchaeota |
| Superphylum: | Asgard Katarzyna Zaremba-Niedzwiedzka, et al. 2017 |
| Phyla | |
|
see text | |
| |
| Synonyms | |
| |
Asgard or Asgardarchaeota[2] is a proposed superphylum consisting of a group of archaea that contain eukaryotic signature proteins.[3] It appears that the eukaryotes, the domain that contains the animals, plants, and fungi, emerged within the Asgard,[4] in a branch containing the Heimdallarchaeota.[5] This supports the two-domain system of classification over the three-domain system.[6][7]
Discovery and nomenclature
In the summer of 2010, sediments were analysed from a gravity core taken in the rift valley on the Knipovich ridge in the Arctic Ocean, near the Loki's Castle hydrothermal vent site. Specific sediment horizons previously shown to contain high abundances of novel archaeal lineages were subjected to metagenomic analysis.[8][9] In 2015, an Uppsala University-led team proposed the Lokiarchaeota phylum based on phylogenetic analyses using a set of highly conserved protein-coding genes.[10] The group was named for the shape-shifting Norse god Loki, in an allusion to the hydrothermal vent complex from which the first genome sample originated.[11] The Loki of mythology has been described as "a staggeringly complex, confusing, and ambivalent figure who has been the catalyst of countless unresolved scholarly controversies",[12] analogous to the role of Lokiarchaeota in the debates about the origin of eukaryotes.[10][13]
In 2016, a University of Texas-led team discovered Thorarchaeota from samples taken from the White Oak River in North Carolina, named in reference to Thor, another Norse god.[14] Samples from Loki's Castle, Yellowstone National Park, Aarhus Bay, an aquifer near the Colorado River, New Zealand's Radiata Pool, hydrothermal vents near Taketomi Island, Japan, and the White Oak River estuary in the United States contained Odinarchaeota and Heimdallarchaeota;[3] following the Norse deity naming convention, these groups were named for Odin and Heimdallr respectively. Researchers therefore named the superphylum containing these microbes "Asgard", after the home of the gods in Norse mythology.[3] Two Lokiarchaeota specimens have been cultured, enabling a detailed insight into their morphology.[15]
Description
Proteins
Asgard members encode many eukaryotic signature proteins, including novel GTPases, membrane-remodelling proteins like ESCRT and SNF7, a ubiquitin modifier system, and N-glycosylation pathway homologs.[3]
Asgard archaeons have a regulated actin cytoskeleton, and the profilins and gelsolins they use can interact with eukaryotic actins.[16][17] In addition, Asgard archaea tubulin from hydrothermal-living Odinarchaeota (OdinTubulin) was identified as a genuine tubulin. OdinTubulin forms protomers and protofilaments most similar to eukaryotic microtubules, yet assembles into ring systems more similar to FtsZ, indicating that OdinTubulin may represent an evolution intermediate between FtsZ and microtubule-forming tubulins.[18] They also seem to form vesicles under cryogenic electron microscopy. Some may have a PKD domain S-layer.[19] They also share the three-way ES39 expansion in LSU rRNA with eukaryotes.[20]
Metabolism
-
Metabolic pathways of Asgard archaea, varying by phyla[21]
-
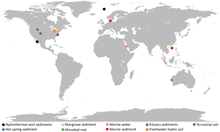
Metabolic pathways of Asgard archaea, varying by environment[21]
Asgard archaea are generally obligate anaerobes, though Kariarchaeota, Gerdarchaeota and Hodarchaeota may be facultative aerobes.[22] They have a Wood–Ljungdahl pathway and perform glycolysis. Members can be autotrophs, heterotrophs, or phototrophs using heliorhodopsin.[21] One member, Candidatus Prometheoarchaeum syntrophicum, is syntrophic with a sulfur-reducing proteobacteria and a methanogenic archaea.[19]
The RuBisCO they have is not carbon-fixing, but likely used for nucleoside salvaging.[21]
Ecology
Asgard are widely distributed around the world, both geographically and by habitat. Many of the known clades are restricted to sediments, whereas Lokiarchaeota, Thorarchaeota and another clade occupy many different habitats. Salinity and depth are important ecological drivers for most Asgard archaea. Other habitats include the bodies of animals, the rhizosphere of plants, non-saline sediments and soils, the sea surface, and freshwater. In addition, Asgard are associated with several other microorganisms.[23]
Eukaryote-like features in subdivisions
The phylum Heimdallarchaeota was found in 2017 to have N-terminal core histone tails, a feature previously thought to be exclusively eukaryotic. Two other archaeal phyla, both outside of Asgard, were found to also have tails in 2018.[24]
In January 2020, scientists found Candidatus Prometheoarchaeum syntrophicum, a member of the Lokiarcheota, engaging in cross-feeding with two bacterial species. Drawing an analogy to symbiogenesis, they consider this relationship a possible link between the simple prokaryotic microorganisms and the complex eukaryotic microorganisms occurring approximately two billion years ago.[25][19]
Classification
The phylogenetic relationships of the Asgard archaea are still under discussion.
| Williams et al. 2019,[5] Eme et al. 2017,[4] Liu et al. 2021[26] & Liu et al. 2020[22] | 53 marker proteins based GTDB 07-RS207[27][28][29] | |||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
In 2023, Eme, Tamarit and colleagues reported that the Eukaryota are deep within Asgard, as sister of Hodarcheales within the Heimdallarchaeia.[30]
| Asgard | |
Taxonomy
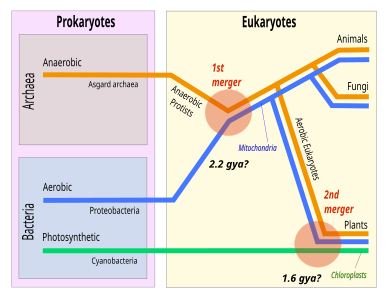
In the depicted scenario, the Eukaryota are deep in the tree of Asgard. A favored scenario is syntrophy, where one organism depends on the feeding of the other. An α-proteobacterium was incorporated to become the mitochondrion.[32] In culture, extant Asgard archaea form various syntrophic dependencies.[33] Gregory Fournier and Anthony Poole have proposed that Asgard is part of "the Eukaryote tree", forming a superphylum they call "Eukaryomorpha" defined by "shared derived characters" (eukaryote signature proteins).[34]
The taxonomy is uncertain and the phylum names are therefore somewhat speculative. The list of phyla is based on the List of Prokaryotic names with Standing in Nomenclature (LPSN)[35] and National Center for Biotechnology Information (NCBI).[36]
- Phylum Baldrarchaeota
- Phylum Borrarchaeota
- Phylum Freyrarchaeota
- Phylum Friggarchaeota
- Phylum Gefionarchaeota
- Phylum Gerdarchaeota
- Phylum Heimdallarchaeota
- Phylum Helarchaeota
- Phylum Hermodarchaeota
- Phylum Hodarchaeota
- Phylum Idunnarchaeota
- Phylum Kariarchaeota
- Phylum Lokiarchaeota
- Phylum Njordarchaeota
- Phylum Odinarchaeota
- Phylum Sifarchaeota
- Phylum Sigynarchaeota
- Phylum Thorarchaeota
- Phylum Tyrarchaeota
- Phylum Wukongarchaeota
Genomic elements
Viruses
Several family-level groups of viruses associated with Asgard archaea have been discovered using metagenomics.[37][38][39] The viruses were assigned to Lokiarchaeia, Thorarchaeia, Odinarchaeia and Helarchaeia hosts using CRISPR spacer matching to the corresponding protospacers within the viral genomes. Two groups of viruses (called 'verdandiviruses') are related to archaeal and bacterial viruses of the class Caudoviricetes, i.e., viruses with icosahedral capsids and helical tails;[37][39] two other distinct groups (called 'skuldviruses') are distantly related to tailless archaeal and bacterial viruses with icosahedral capsids of the realm Varidnaviria;[37][38] and the third group of viruses (called wyrdviruses) is related to archaea-specific viruses with lemon-shaped virus particles (family Halspiviridae).[37][38] The viruses have been identified in deep-sea sediments[37][39] and a terrestrial hot spring of the Yellowstone National Park.[38] All these viruses display very low sequence similarity to other known viruses but are generally related to the previously described prokaryotic viruses,[40] with no meaningful affinity to viruses of eukaryotes.[41][37]
Mobile genetic elements
In addition to viruses, several groups of cryptic mobile genetic elements have been discovered through CRISPR spacer matching to be associated with Asgard archaea of the Lokiarchaeia, Thorarchaeia and Heimdallarchaeia lineages.[37][42] These mobile elements do not encode recognizable viral hallmark proteins and could represent either novel types of viruses or plasmids.
See also
References
- ^ Fournier, G.P.; Poole, A.M. (2018). "A Briefly Argued Case That Asgard Archaea Are Part of the Eukaryote Tree". Frontiers in Microbiology. 9: 1896. doi:10.3389/fmicb.2018.01896. PMC 6104171. PMID 30158917.
- ^ Da Cunha, Violette; Gaia, Morgan; Gadelle, Daniele; et al. (June 2017). "Lokiarchaea are close relatives of Euryarchaeota, not bridging the gap between prokaryotes and eukaryotes". PLOS Genetics. 13 (6): e1006810. doi:10.1371/journal.pgen.1006810. PMC 5484517. PMID 28604769.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ a b c d Zaremba-Niedzwiedzka, Katarzyna; Caceres, Eva F.; Saw, Jimmy H.; et al. (January 2017). "Asgard archaea illuminate the origin of eukaryotic cellular complexity". Nature. 541 (7637): 353–358. Bibcode:2017Natur.541..353Z. doi:10.1038/nature21031. OSTI 1580084. PMID 28077874. S2CID 4458094.
- ^ a b Eme, Laura; Spang, Anja; Lombard, Jonathan; Stairs, Courtney W.; Ettema, Thijs J. G. (November 2017). "Archaea and the origin of eukaryotes". Nature Reviews. Microbiology. 15 (12): 711–723. doi:10.1038/nrmicro.2017.133. PMID 29123225. S2CID 8666687.
- ^ a b Williams, Tom A.; Cox, Cymon J.; Foster, Peter G.; Szöllősi, Gergely J.; Embley, T. Martin (January 2020). "Phylogenomics provides robust support for a two-domains tree of life". Nature Ecology & Evolution. 4 (1): 138–147. doi:10.1038/s41559-019-1040-x. PMC 6942926. PMID 31819234.
- ^ Nobs, Stephanie-Jane; MacLeod, Fraser I.; Wong, Hon Lun; Burns, Brendan P. (May 2022). "Eukarya the chimera: eukaryotes, a secondary innovation of the two domains of life?". Trends in Microbiology. 30 (5): 421–431. doi:10.1016/j.tim.2021.11.003. PMID 34863611. S2CID 244823103.
- ^ Doolittle, W. Ford (February 2020). "Evolution: Two Domains of Life or Three?". Current Biology. 30 (4): R177–R179. doi:10.1016/j.cub.2020.01.010. PMID 32097647.
- ^ Jørgensen, Steffen Leth; Hannisdal, Bjarte; Lanzen, Anders; et al. (October 2012). "Correlating microbial community profiles with geochemical data in highly stratified sediments from the Arctic Mid-Ocean Ridge". PNAS. 109 (42): E2846–E2855. doi:10.1073/pnas.1207574109. PMC 3479504. PMID 23027979.
- ^ Jørgensen, Steffen Leth; Thorseth, Ingunn H.; Pedersen, Rolf B.; et al. (October 4, 2013). "Quantitative and phylogenetic study of the Deep Sea Archaeal Group in sediments of the Arctic mid-ocean spreading ridge". Frontiers in Microbiology. 4: 299. doi:10.3389/fmicb.2013.00299. PMC 3790079. PMID 24109477.
- ^ a b Spang, Anja; Saw, Jimmy H.; Jørgensen, Steffen L.; et al. (May 2015). "Complex archaea that bridge the gap between prokaryotes and eukaryotes". Nature. 521 (7551): 173–179. Bibcode:2015Natur.521..173S. doi:10.1038/nature14447. PMC 4444528. PMID 25945739.
- ^ Yong, Ed. "Break in the Search for the Origin of Complex Life". The Atlantic. Retrieved 2018-03-21.
- ^ von Schnurbein, Stefanie (November 2000). "The Function of Loki in Snorri Sturluson's "Edda"". History of Religions. 40 (2): 109–124. doi:10.1086/463618.
- ^ Spang, Anja; Eme, Laura; Saw, Jimmy H.; et al. (March 2018). "Asgard archaea are the closest prokaryotic relatives of eukaryotes". PLOS Genetics. 14 (3): e1007080. doi:10.1371/journal.pgen.1007080. PMC 5875740. PMID 29596421.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Seitz, Kiley W.; Lazar, Cassandre S.; Hinrichs, Kai-Uwe; et al. (July 2016). "Genomic reconstruction of a novel, deeply branched sediment archaeal phylum with pathways for acetogenesis and sulfur reduction". The ISME Journal. 10 (7): 1696–1705. doi:10.1038/ismej.2015.233. PMC 4918440. PMID 26824177.
- ^ Rodrigues-Oliveira, Thiago; Wollweber, Florian; Ponce-Toledo, Rafael I.; et al. (2023-01-12). "Actin cytoskeleton and complex cell architecture in an Asgard archaeon". Nature. 613 (7943): 332–339. doi:10.1038/s41586-022-05550-y. ISSN 0028-0836. PMC 9834061. PMID 36544020.
- ^ Akıl, Caner; Robinson, Robert C. (October 2018). "Genomes of Asgard archaea encode profilins that regulate actin". Nature. 562 (7727): 439–443. Bibcode:2018Natur.562..439A. doi:10.1038/s41586-018-0548-6. PMID 30283132. S2CID 52917038.
- ^ Akıl, Caner; Tran, Linh T.; Orhant-Prioux, Magali; Baskaran, Yohendran; Manser, Edward; Blanchoin, Laurent; Robinson, Robert C. (August 2020). "Insights into the evolution of regulated actin dynamics via characterization of primitive gelsolin/cofilin proteins from Asgard archaea". Proceedings of the National Academy of Sciences of the United States of America. 117 (33): 19904–19913. bioRxiv 10.1101/768580. doi:10.1073/pnas.2009167117. PMC 7444086. PMID 32747565.
- ^ Akıl, Caner; Ali, Samson; Tran, Linh T.; et al. (March 2022). "Structure and dynamics of Odinarchaeota tubulin and the implications for eukaryotic microtubule evolution". Science Advances. 8 (12): eabm2225. Bibcode:2022SciA....8M2225A. doi:10.1126/sciadv.abm2225. PMC 8956254. PMID 35333570.
- ^ a b c Imachi, Hiroyuki; Nobu, Masaru K.; Nakahara, Nozomi; et al. (January 2020). "Isolation of an archaeon at the prokaryote-eukaryote interface". Nature. 577 (7791): 519–525. Bibcode:2020Natur.577..519I. doi:10.1038/s41586-019-1916-6. PMC 7015854. PMID 31942073.
- ^ Penev, Petar I.; Fakhretaha-Aval, Sara; Patel, Vaishnavi J.; et al. (October 2020). "Supersized Ribosomal RNA Expansion Segments in Asgard Archaea". Genome Biology and Evolution. 12 (10): 1694–1710. doi:10.1093/gbe/evaa170. PMC 7594248. PMID 32785681.
- ^ a b c d MacLeod, Fraser; Kindler, Gareth S.; Wong, Hon Lun; Chen, Ray; Burns, Brendan P. (2019). "Asgard archaea: Diversity, function, and evolutionary implications in a range of microbiomes". AIMS Microbiology. 5 (1): 48–61. doi:10.3934/microbiol.2019.1.48. PMC 6646929. PMID 31384702.
- ^ a b Liu, Yang; Makarova, Kira S.; Huang, Wen-Cong; et al. (2020). "Expanding diversity of Asgard archaea and the elusive ancestry of eukaryotes". bioRxiv. doi:10.1101/2020.10.19.343400. S2CID 225056970.
- ^ Cai, Mingwei; Richter-Heitmann, Tim; Yin, Xiuran; et al. (2021). "Ecological features and global distribution of Asgard archaea". Science of the Total Environment. 758: 143581. doi:10.1016/j.scitotenv.2020.143581. ISSN 0048-9697.
- ^ Henneman, Bram; van Emmerik, Clara; van Ingen, Hugo; Dame, Remus T. (September 2018). "Structure and function of archaeal histones". PLOS Genetics. 14 (9): e1007582. Bibcode:2018BpJ...114..446H. doi:10.1371/journal.pgen.1007582. PMC 6136690. PMID 30212449.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Zimmer, Carl (15 January 2020). "This Strange Microbe May Mark One of Life's Great Leaps - A organism living in ocean muck offers clues to the origins of the complex cells of all animals and plants". The New York Times. Retrieved 16 January 2020.
- ^ Liu, Yang; Makarova, Kira S.; Huang, Wen-Cong; Wolf, Yuri I.; Nikolskaya, Anastasia; Zhang, Xinxu; et al. (May 2021). "Expanded diversity of Asgard archaea and their relationships with eukaryotes". Nature. 593 (7860): 553–557. Bibcode:2021Natur.593..553L. doi:10.1038/s41586-021-03494-3. PMID 33911286. S2CID 233447651.
- ^ "GTDB release 07-RS207". Genome Taxonomy Database. Retrieved 6 December 2021.
- ^ "ar53_r207.sp_label". Genome Taxonomy Database. Retrieved 20 June 2021.
- ^ "Taxon History". Genome Taxonomy Database. Retrieved 6 December 2021.
- ^ Eme, Laura; Tamarit, Daniel; Caceres, Eva F.; et al. (2023-03-09). "Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes". bioRxiv. pp. 2023.03.07.531504. doi:10.1101/2023.03.07.531504v1.
- ^ Latorre, A.; Durban, A.; Moya, A.; Pereto, J. (2011). "The role of symbiosis in eukaryotic evolution". In Gargaud, M.; López-Garcìa, P.; Martin H. (eds.). Origins and Evolution of Life: An astrobiological perspective. Cambridge: Cambridge University Press. pp. 326–339. ISBN 978-0-521-76131-4. Archived from the original on 24 March 2019. Retrieved 27 August 2017.
- ^ López-García, Purificación; Moreira, David (July 2019). "Eukaryogenesis, a syntrophy affair". Nature Microbiology. 4 (7): 1068–1070. doi:10.1038/s41564-019-0495-5. PMC 6684364. PMID 31222170.
- ^ Rodrigues-Oliveira, Thiago; Wollweber, Florian; Ponce-Toledo, Rafael I.; Xu, Jingwei; Rittmann, Simon K.-M. R.; Klingl, Andreas; Pilhofer, Martin; Schleper, Christa (January 2023). "Actin cytoskeleton and complex cell architecture in an Asgard archaeon". Nature. 613 (7943): 332–339. doi:10.1038/s41586-022-05550-y. ISSN 1476-4687. PMC 9834061. PMID 36544020.
- ^ Fournier, Gregory P.; Poole, Anthony M. (2018-08-15). "A Briefly Argued Case That Asgard Archaea Are Part of the Eukaryote Tree". Frontiers in Microbiology. 9. doi:10.3389/fmicb.2018.01896. ISSN 1664-302X.
{{cite journal}}: CS1 maint: unflagged free DOI (link) - ^ Euzéby, J.P. "Superphylum "Asgardarchaeota"". List of Prokaryotic names with Standing in Nomenclature (LPSN). Retrieved 2021-06-27.
- ^ "Asgard group". National Center for Biotechnology Information (NCBI) taxonomy database. Retrieved 2021-03-20.
- ^ a b c d e f g Medvedeva, S.; Sun, J.; Yutin, N.; Koonin, Eugene V.; Nunoura, T.; Rinke, C.; Krupovic, M. (July 2022). "Three families of Asgard archaeal viruses identified in metagenome-assembled genomes". Nature Microbiology. 7 (7): 962–973. doi:10.1038/s41564-022-01144-6. PMID 35760839. S2CID 250091635.
- ^ a b c d Tamarit, D.; Caceres, E.F.; Krupovic, M.; Nijland, R.; Eme, L.; Robinson, N.P.; Ettema, T.J.G. (July 2022). "A closed Candidatus Odinarchaeum chromosome exposes Asgard archaeal viruses". Nature Microbiology. 7 (7): 948–952. doi:10.1038/s41564-022-01122-y. PMC 9246712. PMID 35760836. S2CID 250090798.
- ^ a b c Rambo, I.M.; Langwig, M.V.; Leão, P.; De Anda, V.; Baker, B.J. (July 2022). "Genomes of six viruses that infect Asgard archaea from deep-sea sediments". Nature Microbiology. 7 (7): 953–961. doi:10.1038/s41564-022-01150-8. PMID 35760837.
- ^ Prangishvili, D.; Bamford, D.H.; Forterre, P.; Iranzo, J.; Koonin, Eugene V.; Krupovic, M. (November 2017). "The enigmatic archaeal virosphere". Nature Reviews. Microbiology. 15 (12): 724–739. doi:10.1038/nrmicro.2017.125. PMID 29123227. S2CID 21789564.
- ^ Alarcón-Schumacher, T.; Erdmann, S. (July 2022). "A trove of Asgard archaeal viruses". Nature Microbiology. 7 (7): 931–932. doi:10.1038/s41564-022-01148-2. PMID 35760838. S2CID 250091028.
- ^ Wu, F.; Speth, D.R.; Philosof, A.; et al. (February 2022). "Unique mobile elements and scalable gene flow at the prokaryote-eukaryote boundary revealed by circularized Asgard archaea genomes". Nature Microbiology. 7 (2): 200–212. doi:10.1038/s41564-021-01039-y. PMC 8813620. PMID 35027677.
External links
- Traci Watson: The trickster microbes that are shaking up the tree of life, in: Nature, 14 May 2019

![Metabolic pathways of Asgard archaea, varying by phyla[21]](https://upload.wikimedia.org/wikipedia/commons/thumb/7/7f/Asgard_archaea_Phyla_%28cropped%29.png/634px-Asgard_archaea_Phyla_%28cropped%29.png)
![Metabolic pathways of Asgard archaea, varying by environment[21]](https://upload.wikimedia.org/wikipedia/commons/thumb/a/aa/Asgard_archaea_in_various_environments_%28cropped%29.png/638px-Asgard_archaea_in_various_environments_%28cropped%29.png)

