Xook1kai Choa6aur (talk | contribs) - 'of organisms': DNA may as well come from cell, tissue, organ, organellum (mtDNA ctDNA), population, ecosystem (Archea), secimen, dirt, historical object, ...all of them not being organism |
Adrian J. Hunter (talk | contribs) |
||
| Line 52: | Line 52: | ||
== Large-scale sequencing strategies == |
== Large-scale sequencing strategies == |
||
''[[Shotgun sequencing|shotgun methods]]'' are useful for reading large genomes, but shotgun methods may fail recognize [[sequence repeat]]s often causing omissions or gaps in genome assembly. |
|||
| ⚫ | [[Image:DNA Sequencing gDNA libraries.jpg|thumb|left|Genomic DNA is fragmented into random pieces and cloned as a bacterial library. DNA from individual bacterial clones is sequenced and the sequence is assembled by using overlapping DNA regions.(click to expand)]] Large-scale sequencing aims at sequencing very long DNA pieces, such as whole [[chromosome]]s. Common approaches consist of cutting (with [[restriction enzyme]]s) or shearing (with mechanical forces) large DNA fragments into shorter DNA fragments. The fragmented DNA is [[clone (genetics)|cloned]] into a [[Vector DNA|DNA vector]], and amplified in ''[[Escherichia coli]]''. Short DNA fragments purified from individual bacterial colonies are individually sequenced and [[sequence assembly|assembled electronically]] into one long, contiguous sequence. This method does not require any pre-existing information about the sequence of the DNA and is referred to as ''de novo'' sequencing. Gaps in the assembled sequence may be filled by [[primer walking]]. The different strategies have different tradeoffs in speed and accuracy; ''[[Shotgun sequencing|shotgun methods]]'' are often used for sequencing large genomes, but its assembly is complex and difficult, particularly with [[microsatellites|sequence repeat]]s often causing gaps in genome assembly. |
||
For complex samples, highly repeated sequences, or to preserve samples, DNA may be stored into gnomic library. DNA library method was also popular in early sequencing days when DNA particles were aligned one by one. |
|||
| ⚫ | [[Image:DNA Sequencing gDNA libraries.jpg|thumb|left| |
||
==New sequencing methods== |
==New sequencing methods== |
||
Revision as of 05:46, 30 August 2009
The term DNA sequencing refers to sequencing methods to read genetic information - the order of the nucleotide bases, adenine, guanine, cytosine, and thymine, in a molecule of DNA.
Knowledge of DNA sequences of genes and other regions of the genome has become indispensable for molecular biology and basic research studying biological processes.
The advent of DNA sequencing has significantly accelerated biological research and discovery. The rapid speed of sequencing attained with modern DNA sequencing technology has been instrumental in the sequencing of the human genome, in the Human Genome Project. Related projects, often by scientific collaboration across continents, have generated the complete DNA sequences of many animal, plant, and microbial genomes.

History
RNA sequencing was one of the earliest forms of nucleotide sequencing. The major landmark of RNA sequencing is the sequence of the first complete gene and the complete genome of Bacteriophage MS2, identified and published by Walter Fiers and his coworkers at the University of Ghent (Ghent, Belgium), between 1972[1] and 1976.[2]
Prior to the development of rapid DNA sequencing methods in the early 1970s by Frederick Sanger at the University of Cambridge, in England and Walter Gilbert and Allan Maxam at Harvard,[3][4] a number of laborious methods were used. For instance, in 1973, Gilbert and Maxam reported the sequence of 24 basepairs using a method known as wandering-spot analysis. [5]
The chain-termination method developed by Sanger and coworkers in 1975 soon became the method of choice, owing to its relative ease and reliability.[6][7]
Maxam-Gilbert sequencing
In 1976-1977, Allan Maxam and Walter Gilbert developed a DNA sequencing method based on chemical modification of DNA and subsequent cleavage at specific bases.[3] Although Maxam and Gilbert published their chemical sequencing method two years after the ground-breaking paper of Sanger and Coulson on plus-minus sequencing,[6][8] Maxam-Gilbert sequencing rapidly became more popular, since purified DNA could be used directly, while the initial Sanger method required that each read start be cloned for production of single-stranded DNA. However, with the improvement of the chain-termination method (see below), Maxam-Gilbert sequencing has fallen out of favour due to its technical complexity prohibiting its use in standard molecular biology kits, extensive use of hazardous chemicals, and difficulties with scale-up.
The method requires radioactive labelling at one end and purification of the DNA fragment to be sequenced. Chemical treatment generates breaks at a small proportion of one or two of the four nucleotide bases in each of four reactions (G, A+G, C, C+T). Thus a series of labelled fragments is generated, from the radiolabelled end to the first 'cut' site in each molecule. The fragments in the four reactions are arranged side by side in gel electrophoresis for size separation. To visualize the fragments, the gel is exposed to X-ray film for autoradiography, yielding a series of dark bands each corresponding to a radiolabelled DNA fragment, from which the sequence may be inferred.
Also sometimes known as "chemical sequencing", this method originated in the study of DNA-protein interactions (footprinting), nucleic acid structure and epigenetic modifications to DNA, and within these it still has important applications.
Chain-termination methods

Because the chain-terminator method (or Sanger method after its developer Frederick Sanger) is more efficient and uses fewer toxic chemicals and lower amounts of radioactivity than the method of Maxam and Gilbert, it rapidly became the method of choice. The key principle of the Sanger method was the use of dideoxynucleotide triphosphates (ddNTPs) as DNA chain terminators.
The classical chain-termination method requires a single-stranded DNA template, a DNA primer, a DNA polymerase, radioactively or fluorescently labeled nucleotides, and modified nucleotides that terminate DNA strand elongation. The DNA sample is divided into four separate sequencing reactions, containing all four of the standard deoxynucleotides (dATP, dGTP, dCTP and dTTP) and the DNA polymerase. To each reaction is added only one of the four dideoxynucleotides (ddATP, ddGTP, ddCTP, or ddTTP) which are the chain-terminating nucleotides, lacking a 3'-OH group required for the formation of a phosphodiester bond between two nucleotides, thus terminating DNA strand extension and resulting in DNA fragments of varying length.
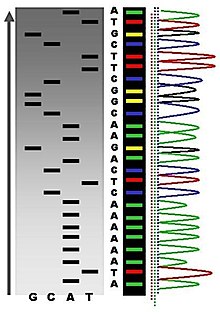
The newly synthesized and labeled DNA fragments are heat denatured, and separated by size (with a resolution of just one nucleotide) by gel electrophoresis on a denaturing polyacrylamide-urea gel with each of the four reactions run in one of four individual lanes (lanes A, T, G, C); the DNA bands are then visualized by autoradiography or UV light, and the DNA sequence can be directly read off the X-ray film or gel image. In the image on the right, X-ray film was exposed to the gel, and the dark bands correspond to DNA fragments of different lengths. A dark band in a lane indicates a DNA fragment that is the result of chain termination after incorporation of a dideoxynucleotide (ddATP, ddGTP, ddCTP, or ddTTP). The relative positions of the different bands among the four lanes are then used to read (from bottom to top) the DNA sequence.

Technical variations of chain-termination sequencing include tagging with nucleotides containing radioactive phosphorus for radiolabelling, or using a primer labeled at the 5’ end with a fluorescent dye. Dye-primer sequencing facilitates reading in an optical system for faster and more economical analysis and automation. The later development by Leroy Hood and coworkers [9][10] of fluorescently labeled ddNTPs and primers set the stage for automated, high-throughput DNA sequencing.
Chain-termination methods have greatly simplified DNA sequencing. For example, chain-termination-based kits are commercially available that contain the reagents needed for sequencing, pre-aliquoted and ready to use. Limitations include non-specific binding of the primer to the DNA, affecting accurate read-out of the DNA sequence, and DNA secondary structures affecting the fidelity of the sequence.
Dye-terminator sequencing

Dye-terminator sequencing utilizes labelling of the chain terminator ddNTPs, which permits sequencing in a single reaction, rather than four reactions as in the labelled-primer method. In dye-terminator sequencing, each of the four dideoxynucleotide chain terminators is labelled with fluorescent dyes, each of which with different wavelengths of fluorescence and emission. Owing to its greater expediency and speed, dye-terminator sequencing is now the mainstay in automated sequencing. Its limitations include dye effects due to differences in the incorporation of the dye-labelled chain terminators into the DNA fragment, resulting in unequal peak heights and shapes in the electronic DNA sequence trace chromatogram after capillary electrophoresis (see figure to the right). This problem has been addressed with the use of modified DNA polymerase enzyme systems and dyes that minimize incorporation variability, as well as methods for eliminating "dye blobs". The dye-terminator sequencing method, along with automated high-throughput DNA sequence analyzers, is now being used for the vast majority of sequencing projects.
Challenges
Common challenges of DNA sequencing include poor quality in the first 15-40 bases of the sequence and deteriorating quality of sequencing traces after 700-900 bases. Base calling software typically gives an estimate of quality to aid in quality trimming.
In cases where DNA fragments are cloned before sequencing, the resulting sequence may contain parts of the cloning vector. In contrast, PCR-based cloning and emerging sequencing technologies based on pyrosequencing often avoid using cloning vectors.
Automation and sample preparation

Automated DNA-sequencing instruments (DNA sequencers) can sequence up to 384 DNA samples in a single batch (run) in up to 24 runs a day. DNA sequencers carry out capillary electrophoresis for size separation, detection and recording of dye fluorescence, and data output as fluorescent peak trace chromatograms. Sequencing reactions by thermocycling, cleanup and re-suspension in a buffer solution before loading onto the sequencer are performed separately. A number of commercial and non-commercial software packages can trim low-quality DNA traces automatically. These programs score the quality of each peak and remove low-quality base peaks (generally located at the ends of the sequence). The accuracy of such algorithms is below visual examination by a human operator, but sufficient for automated processing of large sequence data sets.
Large-scale sequencing strategies
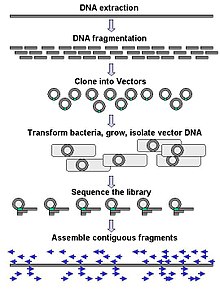
Large-scale sequencing aims at sequencing very long DNA pieces, such as whole chromosomes. Common approaches consist of cutting (with restriction enzymes) or shearing (with mechanical forces) large DNA fragments into shorter DNA fragments. The fragmented DNA is cloned into a DNA vector, and amplified in Escherichia coli. Short DNA fragments purified from individual bacterial colonies are individually sequenced and assembled electronically into one long, contiguous sequence. This method does not require any pre-existing information about the sequence of the DNA and is referred to as de novo sequencing. Gaps in the assembled sequence may be filled by primer walking. The different strategies have different tradeoffs in speed and accuracy; shotgun methods are often used for sequencing large genomes, but its assembly is complex and difficult, particularly with sequence repeats often causing gaps in genome assembly.
New sequencing methods
High-throughput sequencing
The high demand for low-cost sequencing has driven the development of high-throughput sequencing technologies that parallelize the sequencing process, producing thousands or millions of sequences at once.[11][12] High-throughput sequencing technologies are intended to lower the cost of DNA sequencing beyond what is possible with standard dye-terminator methods.
In vitro clonal amplification
Molecular detection methods are not sensitive enough for single molecule sequencing, so most approaches use an in vitro cloning step to amplify individual DNA molecules. Emulsion PCR isolates individual DNA molecules along with primer-coated beads in aqueous droplets within an oil phase. Polymerase chain reaction (PCR) then coats each bead with clonal copies of the DNA molecule followed by immobilization for later sequencing. Emulsion PCR is used in the methods by Marguilis et al. (commercialized by 454 Life Sciences), Shendure and Porreca et al. (also known as "Polony sequencing") and SOLiD sequencing, (developed by Agencourt, now Applied Biosystems).[13][14][15] Another method for in vitro clonal amplification is bridge PCR, where fragments are amplified upon primers attached to a solid surface. The single-molecule method developed by Stephen Quake's laboratory (later commercialized by Helicos) skips this amplification step, directly fixing DNA molecules to a surface.[16]
Parallelized sequencing
DNA molecules are physically bound to a surface, and sequenced in parallel.Sequencing by synthesis, like dye-termination electrophoretic sequencing, uses a DNA polymerase to determine the base sequence. Reversible terminator methods (used by Illumina and Helicos) use reversible versions of dye-terminators, adding one nucleotide at a time, detect fluorescence at each position in real time, by repeated removal of the blocking group to allow polymerization of another nucleotide. Pyrosequencing (used by 454) also uses DNA polymerization, adding one nucleotide species at a time and detecting and quantifying the number of nucleotides added to a given location through the light emitted by the release of attached pyrophosphates.[13][17]
Sequencing by ligation
Sequencing by ligation method uses a DNA ligase to determine the target sequence.[14][15][18] Used in the polony method and in the SOLiD technology, it uses a pool of all possible oligonucleotides of a fixed length, labeled according to the sequenced position. Oligonucleotides are annealed and ligated; the preferential ligation by DNA ligase for matching sequences results in a signal informative of the nucleotide at that position.
Microfluidic Sanger sequencing
In microfluidic Sanger sequencing the entire thermocycling amplification of DNA fragments as well as their separation by electrophoresis is done on a single chip (approximately 100 cm in diameter) thus reducing the reagent usage as well as cost.[citation needed] In some instances researchers [who?] have shown that they can increase the through-put of conventional sequencing through the use of microchips.[citation needed] Research will still need to be done in order to make this use of technology effective.
Other sequencing technologies
Sequencing by hybridization is a non-enzymatic method that uses a DNA microarray. A single pool of DNA whose sequence is to be determined is fluorescently labeled and hybridized to an array containing known sequences. Strong hybridization signals from a given spot on the array identifies its sequence in the DNA being sequenced.[19] Mass spectrometry may be used to determine mass differences between DNA fragments produced in chain-termination reactions.[20]
DNA sequencing methods currently under development include labeling the DNA polymerase,[21] reading the sequence as a DNA strand transits through nanopores,[22][23] and microscopy-based techniques, such as AFM or electron microscopy that are used to identify the positions of individual nucleotides within long DNA fragments (>5,000 bp) by nucleotide labeling with heavier elements (e.g., halogens) for visual detection and recording.[24]
In October 2006, the X Prize Foundation established an initiative to promote the development of full genome sequencing technologies, called the Archon X Prize, intending to award $10 million to "the first Team that can build a device and use it to sequence 100 human genomes within 10 days or less, with an accuracy of no more than one error in every 100,000 bases sequenced, with sequences accurately covering at least 98% of the genome, and at a recurring cost of no more than $10,000 (US) per genome."[25]
Major landmarks in DNA sequencing
- 1953 Discovery of the structure of the DNA double helix.
- 1972 Development of recombinant DNA technology, which permits isolation of defined fragments of DNA; prior to this, the only accessible samples for sequencing were from bacteriophage or virus DNA.
- 1975 The first complete DNA genome to be sequenced is that of bacteriophage φX174
- 1977 Allan Maxam and Walter Gilbert publish "DNA sequencing by chemical degradation".[3] Frederick Sanger, independently, publishes "DNA sequencing by enzymatic synthesis".
- 1980 Frederick Sanger and Walter Gilbert receive the Nobel Prize in Chemistry
- 1984 Medical Research Council scientists decipher the complete DNA sequence of the Epstein-Barr virus, 170 kb.
- 1986 Leroy E. Hood's laboratory at the California Institute of Technology and Smith announce the first semi-automated DNA sequencing machine.
- 1987 Applied Biosystems markets first automated sequencing machine, the model ABI 370.
- 1990 The U.S. National Institutes of Health (NIH) begins large-scale sequencing trials on Mycoplasma capricolum, Escherichia coli, Caenorhabditis elegans, and Saccharomyces cerevisiae (at 75 cents (US)/base).
- 1995 Richard Mathies et al. publish dye-based sequencing.[26]
- 1998 Phil Green and Brent Ewing of the University of Washington publish
“phred”for sequencer data analysis [27].
See also
- Sequencing
- Full genome
- Full genome sequencing
- Genome project
- Single Molecule Real Time Sequencing
- Applied Biosystems
- 454 Life Sciences
- Illumina (company)
- Pacific Biosciences
- Complete Genomics
- Joint Genome Institute
- DNA field-effect transistor
- DNA sequencing theory
References
- ^ Min Jou W, Haegeman G, Ysebaert M, Fiers W (1972). "Nucleotide sequence of the gene coding for the bacteriophage MS2 coat protein". Nature. 237 (5350): 82–8. doi:10.1038/237082a0. PMID 4555447.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Fiers W, Contreras R, Duerinck F; et al. (1976). "Complete nucleotide sequence of bacteriophage MS2 RNA: primary and secondary structure of the replicase gene". Nature. 260 (5551): 500–7. doi:10.1038/260500a0. PMID 1264203.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ a b c Maxam AM, Gilbert W (1977). "A new method for sequencing DNA". Proc. Natl. Acad. Sci. U.S.A. 74 (2): 560–4. doi:10.1073/pnas.74.2.560. PMC 392330. PMID 265521.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Gilbert, W. DNA sequencing and gene structure. Nobel lecture, 8 December 1980.
- ^ Gilbert W, Maxam A (1973). "The nucleotide sequence of the lac operator". Proc. Natl. Acad. Sci. U.S.A. 70 (12): 3581–4. doi:10.1073/pnas.70.12.3581. PMC 427284. PMID 4587255.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ a b Sanger F, Coulson AR (1975). "A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase". J. Mol. Biol. 94 (3): 441–8. doi:10.1016/0022-2836(75)90213-2. PMID 1100841.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Sanger F, Nicklen S, Coulson AR (1977). "DNA sequencing with chain-terminating inhibitors". Proc. Natl. Acad. Sci. U.S.A. 74 (12): 5463–7. doi:10.1073/pnas.74.12.5463. PMC 431765. PMID 271968.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Sanger F. Determination of nucleotide sequences in DNA. Nobel lecture, 8 December 1980.
- ^ Smith LM, Sanders JZ, Kaiser RJ; et al. (1986). "Fluorescence detection in automated DNA sequence analysis". Nature. 321 (6071): 674–9. doi:10.1038/321674a0. PMID 3713851.
We have developed a method for the partial automation of DNA sequence analysis. Fluorescence detection of the DNA fragments is accomplished by means of a fluorophore covalently attached to the oligonucleotide primer used in enzymatic DNA sequence analysis. A different coloured fluorophore is used for each of the reactions specific for the bases A, C, G and T. The reaction mixtures are combined and co-electrophoresed down a single polyacrylamide gel tube, the separated fluorescent bands of DNA are detected near the bottom of the tube, and the sequence information is acquired directly by computer.
{{cite journal}}: Explicit use of et al. in:|author=(help)CS1 maint: multiple names: authors list (link) - ^ Smith LM, Fung S, Hunkapiller MW, Hunkapiller TJ, Hood LE (1985). "The synthesis of oligonucleotides containing an aliphatic amino group at the 5' terminus: synthesis of fluorescent DNA primers for use in DNA sequence analysis". Nucleic Acids Res. 13 (7): 2399–412. doi:10.1093/nar/13.7.2399. PMC 341163. PMID 4000959.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Hall N (2007). "Advanced sequencing technologies and their wider impact in microbiology". J. Exp. Biol. 210 (Pt 9): 1518–25. doi:10.1242/jeb.001370. PMID 17449817.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ Church GM (2006). "Genomes for all". Sci. Am. 294 (1): 46–54. PMID 16468433.
{{cite journal}}: Unknown parameter|month=ignored (help) - ^ a b Margulies M, Egholm M, Altman WE; et al. (2005). "Genome sequencing in microfabricated high-density picolitre reactors". Nature. 437 (7057): 376–80. doi:10.1038/nature03959. PMC 1464427. PMID 16056220.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ a b Shendure, J. (2005). "Accurate Multiplex Polony Sequencing of an Evolved Bacterial Genome". Science. 309: 1728. doi:10.1126/science.1117389.
- ^ a b Applied Biosystems' SOLiD technology
- ^ Braslavsky I, Hebert B, Kartalov E, Quake SR (2003). "Sequence information can be obtained from single DNA molecules". Proc. Natl. Acad. Sci. U.S.A. 100 (7): 3960–4. doi:10.1073/pnas.0230489100. PMC 153030. PMID 12651960.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ M. Ronaghi, S. Karamohamed, B. Pettersson, M. Uhlen, and P. Nyren (1996). "Real-time DNA sequencing using detection of pyrophosphate release". Analytical Biochemistry. 242: 84–9. doi:10.1006/abio.1996.0432.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ US patent 5750341, Macevicz SC, "DNA sequencing by parallel oligonucleotide extensions", issued 1995-04-17
- ^ Hanna GJ, Johnson VA, Kuritzkes DR; et al. (2000). "Comparison of sequencing by hybridization and cycle sequencing for genotyping of human immunodeficiency virus type 1 reverse transcriptase". J. Clin. Microbiol. 38 (7): 2715–21. PMC 87006. PMID 10878069.
{{cite journal}}: Explicit use of et al. in:|author=(help); Unknown parameter|day=ignored (help); Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ J.R. Edwards, H.Ruparel, and J. Ju (2005). "Mass-spectrometry DNA sequencing". Mutation Research. 573 (1–2): 3–12.
{{cite journal}}: CS1 maint: multiple names: authors list (link) - ^ VisiGen Biotechnologies Inc. - Technology Overview
- ^ The Harvard Nanopore Group
- ^ "Nanopore Sequencing Could Slash DNA Analysis Costs".
- ^ US patent 20060029957, ZS Genetics, "Systems and methods of analyzing nucleic acid polymers and related components", issued 2005-07-14
- ^ "PRIZE Overview: Archon X PRIZE for Genomics"
- ^ Ju J, Ruan C, Fuller CW, Glazer AN, Mathies RA (1995). "Fluorescence energy transfer dye-labeled primers for DNA sequencing and analysis". Proc. Natl. Acad. Sci. U.S.A. 92 (10): 4347–51. doi:10.1073/pnas.92.10.4347. PMC 41941. PMID 7753809.
{{cite journal}}: Unknown parameter|month=ignored (help)CS1 maint: multiple names: authors list (link) - ^ Ewing B, Green P (1998). "Base-calling of automated sequencer traces using phred. II. Error probabilities". Genome Res. 8 (3): 186–94. PMID 9521922.
{{cite journal}}: Unknown parameter|month=ignored (help)
External links
- Disruptive Gene Sequencing technology - Single Molecule Real Time (SMRT) sequencing
